Microsatellite DNA
One type of genetic data that was extracted from the Gouldian finch feathers sent was microsatellite DNA. These are commonly used in all sorts of genetic studies, such as: pinpointing genes responsible for certain traits or genetic diseases, estimating paternity and maternity, and characterising gene-flow between regions. The latter two types of analyses were conducted using microsatellite DNA on Gouldian finches in the wild (Bolton et al 2016; Bolton et al 2017).
The reason microsatellites have been used for so many genetic studies for so long is because they are incredibly variable markers - they have lots of alleles! This makes it easy to distinguish between individuals, espescially when some alleles are very rare. In the Australian domesticated Gouldian finches is locus (gene) named Ego49, with 43 alleles across all of Australia.
These markers are short sequences of DNA repeated multiple times (e.g. ATG 15 times, or 20 times). There are many of these repeated sequences throughout the genome, but we can select a certain region to focus on using a method known as polymerase chain reaction (PCR). Using special short sequences that are designed to bind to adjacent areas of DNA, PCR hijacks the usual DNA replication enzyme machinery and just replicates a particular genetic region over and over again. This allows us to see a particular region in preference over the remainder of the genome.
Microsatellite markers are measured based on their weight. A microsatellite with 15 repeats of ATG is going to be lighter than a microsatellite with 20 repeats. Figure 1 shows an example of the raw microsatellite data from the wild population. The data returned to you is simply a number that represents the weights of each allele extracted from this program, with an allele for each chromosome.
In your instructions document accompanying your data, the original papers that discovered the particular microsatellite locus are referenced.
The reason microsatellites have been used for so many genetic studies for so long is because they are incredibly variable markers - they have lots of alleles! This makes it easy to distinguish between individuals, espescially when some alleles are very rare. In the Australian domesticated Gouldian finches is locus (gene) named Ego49, with 43 alleles across all of Australia.
These markers are short sequences of DNA repeated multiple times (e.g. ATG 15 times, or 20 times). There are many of these repeated sequences throughout the genome, but we can select a certain region to focus on using a method known as polymerase chain reaction (PCR). Using special short sequences that are designed to bind to adjacent areas of DNA, PCR hijacks the usual DNA replication enzyme machinery and just replicates a particular genetic region over and over again. This allows us to see a particular region in preference over the remainder of the genome.
Microsatellite markers are measured based on their weight. A microsatellite with 15 repeats of ATG is going to be lighter than a microsatellite with 20 repeats. Figure 1 shows an example of the raw microsatellite data from the wild population. The data returned to you is simply a number that represents the weights of each allele extracted from this program, with an allele for each chromosome.
In your instructions document accompanying your data, the original papers that discovered the particular microsatellite locus are referenced.
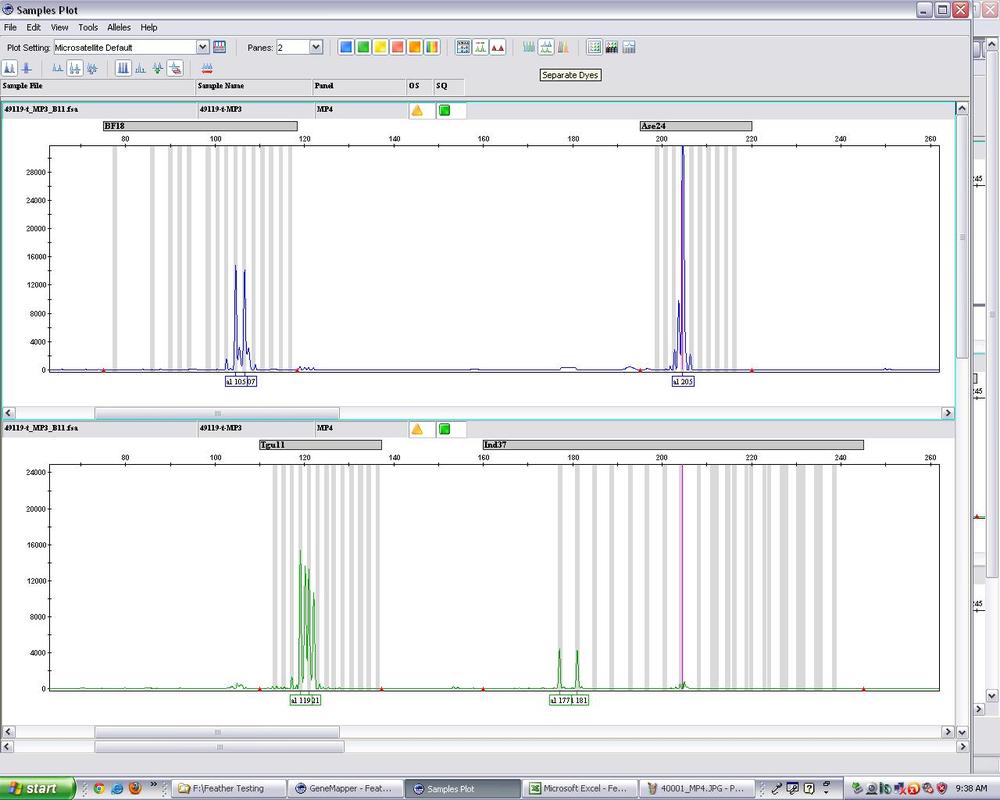
Figure 1: An example of some microsatellite loci from a wild Gouldian finch. Many loci can be amplified in the same reaction, and if they are similar weights, they are given a different dye so that we can distinguish them.
In the figure above locus BF18 (the grey bar above the blue peaks on the top left) is a similar overall size to locus Tgu11 in green.
Locus BF18 is a heterozygote for alleles with sizes 105 and 107, while Ase24 is a homozygote for a single allee at size 205.
Mitochondrial DNA
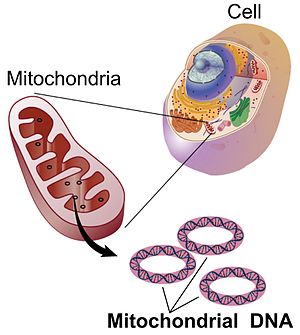 Figure 2. Diagram of the a mitochondrion in a cell, with the circular DNA.
Image from Wikipedia:
https://en.wikipedia.org/wiki/Mitochondrial_DNA
Figure 2. Diagram of the a mitochondrion in a cell, with the circular DNA.
Image from Wikipedia:
https://en.wikipedia.org/wiki/Mitochondrial_DNA
The mitochondria are organelles in the cells of all birds (and humans, and many other organisms too). They have a genome that is inherited separately from the chromosomes in the nucleus, and is only transmitted via the mother in her eggs. There are no chromosomes and the mitochondrial DNA is in a circle (Figure 2). Individuals do not have alleles inherited from mother and father, just the mother.
This marker is also commonly used in genetic studies. Here we have amplified a particular sub-region of the mitochondrial DNA known as the mitochondrial control region. The method we used for the Gouldian finches was developed originally in the Long-tailed finch (Rollins et al 2012).
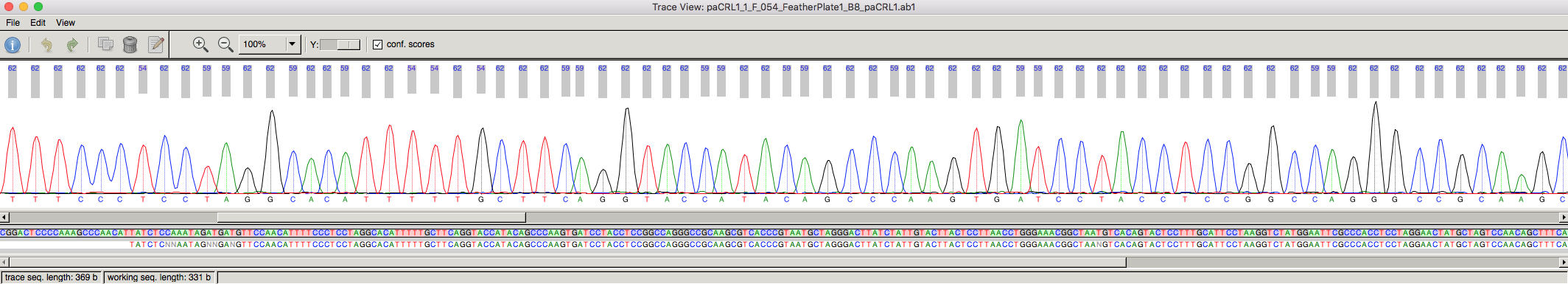
The data output from mitochondrial DNA is different from microsatellites. Here, each base (A, T, G or C) is read by the sequencing machine as a different coloured dye, outputting a sequence. An example of the raw mitochondrial data is provided in Figure 3.
This marker is also commonly used in genetic studies. Here we have amplified a particular sub-region of the mitochondrial DNA known as the mitochondrial control region. The method we used for the Gouldian finches was developed originally in the Long-tailed finch (Rollins et al 2012).
The data output from mitochondrial DNA is different from microsatellites. Here, each base (A, T, G or C) is read by the sequencing machine as a different coloured dye, outputting a sequence. An example of the raw mitochondrial data is provided in Figure 3.
References
Bolton, P.E., West, A.J, Cardili, A.P.A, Clarke, J.A., Maute, K.L., Legge, S., Brazill-Boast, J., Griffith, S.C., & Rollins, L.A. 2016. Three molecular markers show no evidence of population genetic structure in the Gouldian finch (Erythrura gouldiae). Plos One. DOI:10.1371/journal.pone.0167723
Bolton, P.E., Rollins, L.A., Brazill-Boast, J., Kim, K-W, Burke, T., & Griffith, S.C. 2017. The colour of paternity: extra-pair paternity in the wild Gouldian finch does not appear to be driven by genetic incompatibility between morphs. Journal of Evolutionary Biology. 30: 174-190 DOI: 10.1111/jeb.12997
Rollins, L. A., Svedin, N., Pryke, S. R., & Griffith, S. C. 2012. The role of the Ord Arid Intrusion in the historical and contemporary genetic division of long-tailed finch subspecies in northern Australia. Ecology and Evolution. 2: 1208–19. https://doi.org/10.1002/ece3.259
Bolton, P.E., Rollins, L.A., Brazill-Boast, J., Kim, K-W, Burke, T., & Griffith, S.C. 2017. The colour of paternity: extra-pair paternity in the wild Gouldian finch does not appear to be driven by genetic incompatibility between morphs. Journal of Evolutionary Biology. 30: 174-190 DOI: 10.1111/jeb.12997
Rollins, L. A., Svedin, N., Pryke, S. R., & Griffith, S. C. 2012. The role of the Ord Arid Intrusion in the historical and contemporary genetic division of long-tailed finch subspecies in northern Australia. Ecology and Evolution. 2: 1208–19. https://doi.org/10.1002/ece3.259